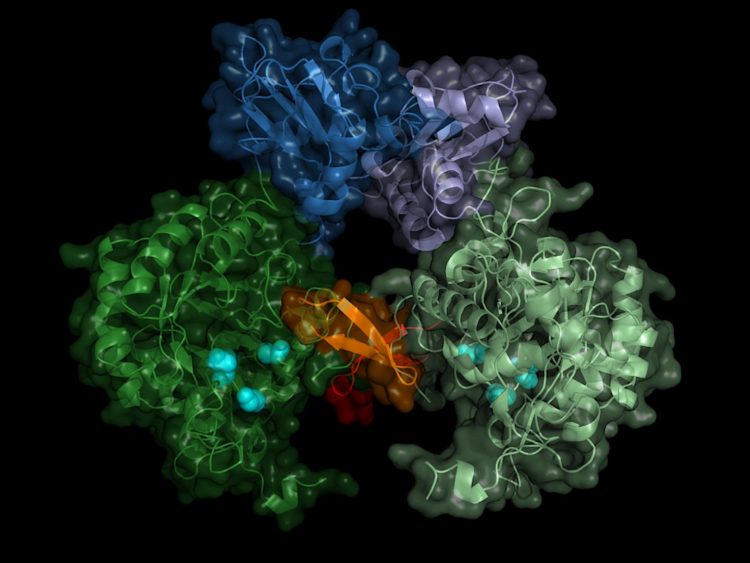
Nuclear Metabolic Fingerprints: How Hidden Enzymes on DNA Could Revolutionize Cancer Treatment
A groundbreaking study published in Nature Communications reveals that over 200 metabolic enzymes are directly attached to human DNA inside the cell nucleus, forming unique 'nuclear metabolic fingerprints' that vary between tissues and cancers. This discovery challenges the traditional separation between metabolism and gene regulation, showing that these enzymes gather around damaged DNA to assist with repair. The research suggests these nuclear enzymes may influence how cancers grow and respond to treatment, potentially explaining why tumors with similar mutations respond differently to therapies. This hidden nuclear metabolism opens new avenues for cancer diagnosis and treatment strategies.
For decades, cellular biology textbooks have presented a clear division of labor: the nucleus houses and regulates our genetic material, while metabolism—the complex network of chemical reactions that sustains life—operates primarily in the cytoplasm and mitochondria. A revolutionary study published in Nature Communications shatters this long-held paradigm. Researchers from the Center for Genomic Regulation have discovered that hundreds of metabolic enzymes are not confined to their traditional compartments but are actively attached to human DNA within the nucleus itself. This hidden network, which varies uniquely between different tissues and cancers, forms what scientists are calling a "nuclear metabolic fingerprint," revealing an unexpected and profound link between cellular energy production and genetic control that could fundamentally alter our understanding of cancer biology and treatment.
Uncovering a Hidden Nuclear World
The discovery emerged from a systematic effort to profile proteins physically bound to chromatin—the complex of DNA and proteins that packages genetic material. Using a specialized isolation technique, the research team analyzed 44 cancer cell lines and 10 healthy cell types from various tissues. The results were astonishing. Approximately 7% of all proteins attached to chromatin were identified as metabolic enzymes, totaling more than 200 different enzymes. Many of these enzymes, such as those involved in oxidative phosphorylation—the primary energy-producing pathway—were traditionally thought to reside exclusively in mitochondria. Their presence on DNA suggests the nucleus operates its own distinct "mini metabolism," a concept that redefines our view of nuclear function.
The Nuclear Metabolic Fingerprint
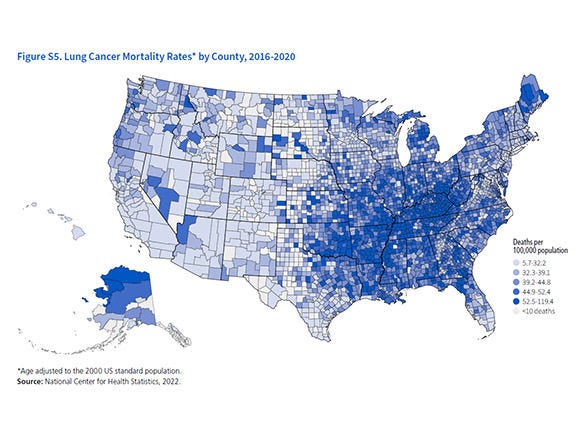
One of the most significant findings is that the pattern of these nuclear-bound enzymes is not random but forms a unique signature for each cell type. The study, as detailed in the ScienceDaily report, revealed that breast cancer cells, for instance, show high nuclear levels of energy-producing oxidative phosphorylation enzymes, while these same enzymes are largely absent in the nuclei of lung cancer cells. This tissue-specific signature, or "fingerprint," was confirmed not only in cell lines but also in actual patient tumor samples. Dr. Sara Sdelci, corresponding author of the study, emphasizes that this variation may directly shape how cancer cells respond to stress and treatment, representing "an entirely new world to explore."
Direct Role in DNA Repair and Genome Stability
Beyond mere presence, the research provides compelling evidence for the functional role of these nuclear enzymes. Experiments demonstrated that a specific group of enzymes responsible for synthesizing molecules essential for DNA building blocks rapidly congregate around sites of DNA damage. This strategic localization suggests they play a direct, active role in repairing genomic injuries—a critical process for cell survival. Furthermore, the function of an enzyme was found to depend entirely on its cellular location. The enzyme IMPDH2, when forced to remain in the nucleus, contributed to maintaining genome stability. When confined to the cytoplasm, it influenced entirely different cellular pathways. This spatial control of enzyme function adds a new layer of complexity to cellular regulation.
Implications for Cancer Therapy and Future Research
The discovery of nuclear metabolism has immediate and profound implications for oncology. It provides a potential mechanistic explanation for a long-observed clinical puzzle: why tumors originating in different tissues, even when they carry identical genetic mutations, often respond very differently to chemotherapy, radiotherapy, or targeted drugs. If metabolic processes and DNA repair are intimately connected within the nucleus, then therapies targeting one system will inevitably affect the other. This interconnectedness could be exploited to develop new combination therapies or to predict treatment resistance. Mapping these nuclear metabolic fingerprints could also lead to new biomarkers for cancer diagnosis and prognosis.
However, the research opens as many questions as it answers. Scientists must now determine whether all these nuclear enzymes are catalytically active and decipher the specific role of each one. A major mystery is how these often-large enzymes bypass the nuclear pore complex, which typically restricts the passage of bulky molecules. Unraveling this transport mechanism could reveal new therapeutic targets. As first author Dr. Savvas Kourtis notes, "We've been treating metabolism and genome regulation as two separate universes, but our work suggests they're talking to each other, and cancer cells might be exploiting these conversations to survive." This dialogue between metabolism and the genome, once hidden, now stands as a promising frontier for understanding and conquering cancer.