How Phage-Pathogen Coevolution Shapes Cholera Outbreaks
New research reveals how the dynamic arms race between cholera-causing bacteria and their viral predators directly influences epidemic patterns. Through clinical surveillance in Bangladesh, scientists captured the real-time evolution of Vibrio cholerae as it acquired a novel anti-phage defense element called PLE11, which initiated a selective sweep coinciding with the largest cholera outbreak in recent records. This study provides the first direct molecular evidence that phage predation drives the selection of epidemic strains, offering crucial insights into how viral pressure shapes pathogen evolution in human disease contexts.
In the complex world of infectious diseases, the relationship between pathogens and their viral predators plays a crucial but often overlooked role in shaping epidemics. Recent research published in Nature reveals how the dynamic coevolution between cholera-causing bacteria and their bacteriophage predators directly influences outbreak patterns and strain selection. This groundbreaking study provides the first direct molecular evidence that phage predation drives the evolution of epidemic Vibrio cholerae strains, offering new insights into the factors that determine which bacterial variants dominate during outbreaks.

The Cholera-Phage Arms Race
Cholera, caused by the bacterium Vibrio cholerae, remains a significant global health threat, particularly in regions like Bangladesh where the disease is endemic. For years, researchers have observed that higher ratios of the lytic phage ICP1 in patient stool correlate with reduced disease severity, suggesting that phage predation might limit cholera outbreaks. However, direct molecular evidence linking phage pressure to the selection of specific epidemic strains has been lacking until now.
The study focused on a family of mobile genetic elements called phage-inducible chromosomal island-like elements (PLEs) that V. cholerae uses to defend against ICP1 predation. These PLEs have evolved intricate mechanisms to disrupt the phage life cycle while exploiting phage machinery for their own transmission. The research team conducted high-resolution clinical surveillance in Bangladesh between October 2019 and June 2022, collecting and analyzing 516 stool samples from patients with suspected cholera.
Capturing a Selective Sweep in Real Time
During their surveillance period, researchers made a critical discovery: the acquisition of a previously unknown PLE variant called PLE11 by the dominant V. cholerae lineage in Bangladesh. This acquisition initiated a selective sweep that coincided with the largest cholera outbreak in recent records, during which the icddr,b Dhaka hospital treated more than 42,000 patients.

PLE11 showed potent anti-phage activity against cocirculating ICP1 strains, explaining its rapid and dominating emergence. Within nine months of its initial detection, PLE11 was present in 91% of V. cholerae isolates. The researchers identified PLE11-encoded Rta as the defense protein responsible for this protection and provided evidence that Rta restricts phage tail assembly, rendering phages incapable of propagation.
Molecular Mechanisms of Defense and Counter-Defense
The study revealed that PLE11 uses a unique strategy to protect V. cholerae populations. Unlike previously characterized PLE variants, PLE11 maintains phage defense even when its genome is targeted for degradation by phage counter-defenses like CRISPR-Cas systems. This is achieved through Rta, which interferes with phage tail assembly independently of the fate of the PLE genome.
Through experimental evolution, researchers predicted how ICP1 would counteradapt to overcome PLE11-mediated defense. Their predictions were validated when continued surveillance documented the emergence of ICP1 strains with CRISPR-Cas systems and specific mutations in their tape measure protein (TMP) that allowed them to evade Rta-mediated restriction. This represented a convergent evolutionary outcome between laboratory predictions and natural selection.
Chimeric Tail Assembly: A Novel Satellite Strategy
One of the most surprising findings was how PLEs balance their dependence on ICP1 tail proteins for horizontal transmission with the restriction of phage tail assembly by Rta. The researchers discovered that PLEs construct chimeric tails composed of both mobile genetic element-encoded and phage-encoded proteins to ensure their transmission.
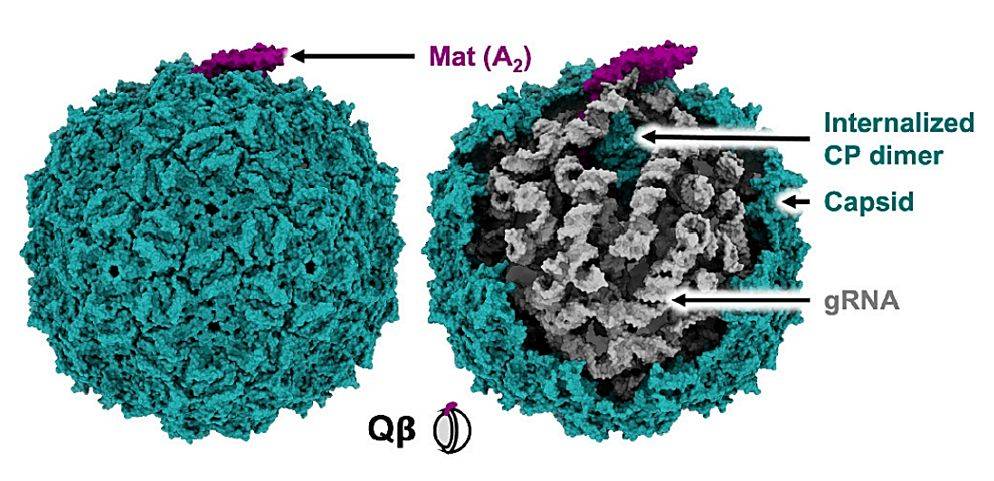
Mass spectrometry and transmission electron microscopy revealed that PLE11 virions incorporate PLE-encoded tape measure proteins and baseplate hub proteins while excluding their phage-encoded counterparts. This represents the first example of phage satellites manipulating viral tail assembly to incorporate both satellite-encoded and phage-encoded structural components into functional chimeric tails.
Implications for Public Health and Surveillance
The findings have significant implications for understanding and responding to cholera epidemics. The study provides strong evidence that predation by lytic phages contributes to the selection of pandemic V. cholerae strains. The acquisition of PLE11 enabled V. cholerae strains to resist contemporaneous phages, likely facilitating their proliferation during the devastating 2022 outbreak.
This research underscores the importance of sustained genomic and phenotypic monitoring of both phages and bacteria to track the emergence and spread of pandemic V. cholerae strains. Beyond single-nucleotide polymorphism-based phylogenetic analyses, examining variable accessory gene content like PLEs is crucial for understanding the success of emerging lineages and their potential to seed global cholera epidemics.
Future Directions and Applications
The study highlights several important directions for future research and public health applications. First, deviations from expected patterns of phage susceptibility could inform public health measures, with phage-resistant lineages prioritized as variants of concern. Second, understanding the coevolution of phages and their bacterial hosts could lead to new therapeutic approaches that leverage natural phage predation dynamics.
Finally, the research demonstrates the value of combining clinical surveillance with mechanistic insight to understand how emerging pathogen lineages successfully spread. As cholera continues to threaten vulnerable populations worldwide, such integrated approaches will be essential for developing effective prevention and control strategies.