Optovolution: Using Light to Engineer Dynamic Proteins with Computational Abilities
Researchers at EPFL have developed a groundbreaking method called optovolution that uses light to guide the evolution of proteins with complex, dynamic behaviors. By engineering yeast cells so their survival depends on proteins switching states at precise times, scientists can rapidly select variants that function like biological switches, sensors, and even simple computers. This technique overcomes limitations of traditional directed evolution and opens new possibilities for synthetic biology, biotechnology, and fundamental research into how proteins evolve complex functionalities.
In the quest to engineer biological systems with unprecedented precision, scientists have developed a revolutionary approach that harnesses light to guide protein evolution. This technique, called optovolution, represents a significant leap forward in synthetic biology by enabling the creation of proteins that can switch, sense, and compute—capabilities that mirror the dynamic behaviors found in natural biological systems. Unlike traditional methods that favor static, always-on proteins, optovolution selects for variants that can change states in response to environmental cues, opening new frontiers in biotechnology and cellular engineering.

The Limitations of Traditional Directed Evolution
Directed evolution has been a powerful tool in biotechnology for decades, allowing scientists to improve enzymes, antibodies, and other proteins for applications ranging from medicine to industrial manufacturing. However, this approach has a fundamental limitation: it typically applies constant selection pressure that rewards proteins that remain highly active at all times. As noted in research from EPFL's Laboratory of the Physics of Biological Systems, this static selection pressure fails to capture the dynamic nature of real biological systems, where proteins often serve as signals, molecular switches, or logic gates that must change states as conditions shift.
The problem with traditional methods becomes apparent when trying to evolve proteins with complex multi-state behaviors. When evolution experiments only reward a single state, other necessary states can degrade over time, causing proteins to lose their ability to switch properly. This limitation has made it difficult to create proteins that can perform the sophisticated computational tasks that occur naturally in living cells, where timing and state transitions are just as important as signal strength.
How Optovolution Works: A Light-Guided Approach
Optovolution represents a paradigm shift in protein engineering by using light to steer evolution toward dynamic functionalities. The method, developed by researchers led by Sahand Jamal Rahi at EPFL, employs optogenetics—a technique that uses light to activate or deactivate genes—to create a selection system that rewards proper timing and state transitions rather than constant activity.
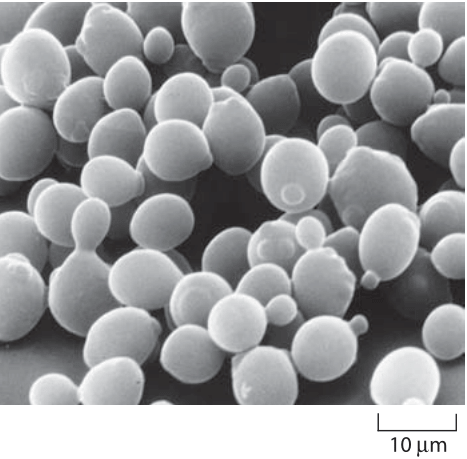

The researchers engineered budding yeast cells (Saccharomyces cerevisiae) so that cell division depended entirely on the behavior of the protein being evolved. They connected the protein's output signal to a regulator that controls the cell cycle—a regulator that is essential during one stage but becomes toxic during another. This clever design meant that if the protein remained either on or off for too long, the yeast cell would stall or die. Only cells containing proteins that switched at precisely the correct time could continue to divide and pass on their genetic material.
By delivering timed pulses of light, the researchers forced the protein to alternate between active and inactive states. Each yeast cell cycle lasts approximately 90 minutes, creating a rapid pass-or-fail test of whether the protein switched at the correct moment. Proteins that performed best allowed their host cells to survive and reproduce, while poorly switching variants were automatically eliminated. This system enabled optovolution to select for proteins with superior dynamic behavior without requiring manual screening or repeated adjustments.
Breakthrough Applications and Protein Variants
The optovolution technique has already yielded several remarkable achievements in protein engineering. Researchers first improved a commonly used light-controlled transcription factor, generating 19 new variants with enhanced properties. Some variants showed greater sensitivity to light, while others exhibited reduced activity in darkness—a crucial feature for minimizing background noise in biological systems. Perhaps most impressively, the team evolved proteins that respond to green light rather than only blue light, overcoming a long-standing challenge in optogenetics.
Engineering proteins that respond to warmer colors like green has been considered extremely difficult due to how these proteins absorb light. The success of optovolution in this area demonstrates its power to achieve results that have eluded traditional methods. Additionally, researchers evolved a red light optogenetic system that no longer requires an added chemical cofactor. Evolution produced an unexpected mutation that disabled a normal yeast transport protein, allowing the system to use light-sensitive molecules already present inside the cell. This simplification makes the system easier to use in experimental settings and represents a more elegant biological solution.

Proteins as Biological Computers
Perhaps the most exciting demonstration of optovolution's potential is its ability to evolve proteins that function like tiny biological computers. The researchers successfully evolved a transcription factor that activates genes only when two different inputs appear simultaneously—one light signal and one chemical signal. This protein behaves like a biological logic gate, making yes-or-no decisions based on multiple environmental cues.
This capability represents a significant advancement toward creating sophisticated cellular circuits that can process information and make decisions autonomously. Such proteins could enable the development of smart therapeutic cells that respond only when specific disease markers are present, or industrial microorganisms that optimize their metabolic pathways based on multiple environmental factors. The ability to evolve proteins with computational functionalities brings synthetic biology closer to creating truly intelligent biological systems.
Future Implications and Applications
Optovolution opens numerous possibilities across biotechnology, synthetic biology, and fundamental research. The technique may help scientists design smarter cellular circuits for therapeutic applications, create optogenetic tools that respond independently to different colors of light, and better understand how complex protein behaviors arise through evolution. By enabling dynamic behaviors to evolve continuously within living cells, optovolution offers a more natural approach to protein engineering that mirrors how evolution operates in nature.
The research, published in the journal Cell, represents a collaborative effort involving multiple institutions including EPFL's Laboratory of Protein and Cell Engineering, the University of Bayreuth, and Lausanne University Hospital (CHUV). As the technique continues to develop, it promises to accelerate progress in areas ranging from biomedical engineering to sustainable biomanufacturing, ultimately helping scientists harness the full potential of biological systems for human benefit.